Researchers at Rice University and the University of Pennsylvania have refined a DNA base-editing technology to make it more precise, a step that could improve prospects for treating some patients with cystic fibrosis and increase the safety and efficacy of therapies for a range of genetic disorders.

Unlike infectious diseases, many of which respond to the same treatments — like antibiotics that neutralize multiple types of bacteria — genetic diseases often require extremely specific therapies, which can vary from patient to patient, even for the same disease.
For example, more than 1,000 different mutations can cause cystic fibrosis, so in order to treat it, researchers need to develop a suite of tools rather than a single therapy. But even when scientists know exactly which DNA letter they want to change, today’s gene-editing technologies can unintentionally alter nearby letters as well, introducing “bystander” mutations that raise safety concerns.
“We were able to introduce specific cystic fibrosis mutations into human epithelial cells relevant to the disease to generate cystic fibrosis cell models for better understanding the disease,” said Gang Bao, the Foyt Family Professor of Bioengineering at Rice and co-senior author of the study, published in Molecular Therapy. “We were also able to reverse those mutations and show improved cellular functions using the same editor, demonstrating the level of pinpoint gene-editing control this technology now offers and showing the potential of using base editing for treating cystic fibrosis.”
One common cause of genetic diseases, including cystic fibrosis, is the accidental substitution of one nucleotide base — a single “letter” in the genetic code — for another.
“In some cases, the letter should be a T,” said Tyler C. Daniel, a Penn doctoral candidate in chemical and biomolecular engineering and co-first author of the new paper, referring to thymine, one of the four bases in human DNA, along with adenine (A), guanine (G) and cytosine (C). “Instead, it’s a C, which can impair or completely abolish the function of the gene, leading to disease.”
While it’s already possible to change the C to a T, including with a base-pair editor the same researchers devised in 2020, and even to selectively modify just one of two adjacent Cs, problems arise when multiple pairs of cytosines appear close together, in “CC … CC” patterns, separated by just a few other base pairs.
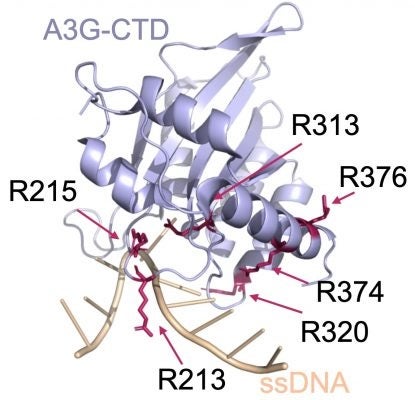
The challenge is hardly theoretical: Among the tens of thousands of known disease-causing C-to-T and T-to-C mutations this type of base-pair editor can address, three-quarters involve multiple cytosine pairs clustered together.
“The fact that different mutations require distinct corrective tools highlights the importance of precision medicine,” said Xue “Sherry” Gao, the Presidential Penn Compact Associate Professor in Chemical and Biomolecular Engineering and Bioengineering at Penn and co-senior author on the study. “We can already identify and replace a particular letter in a specific word. How do we change only that one letter without accidentally altering the letters next to it?”
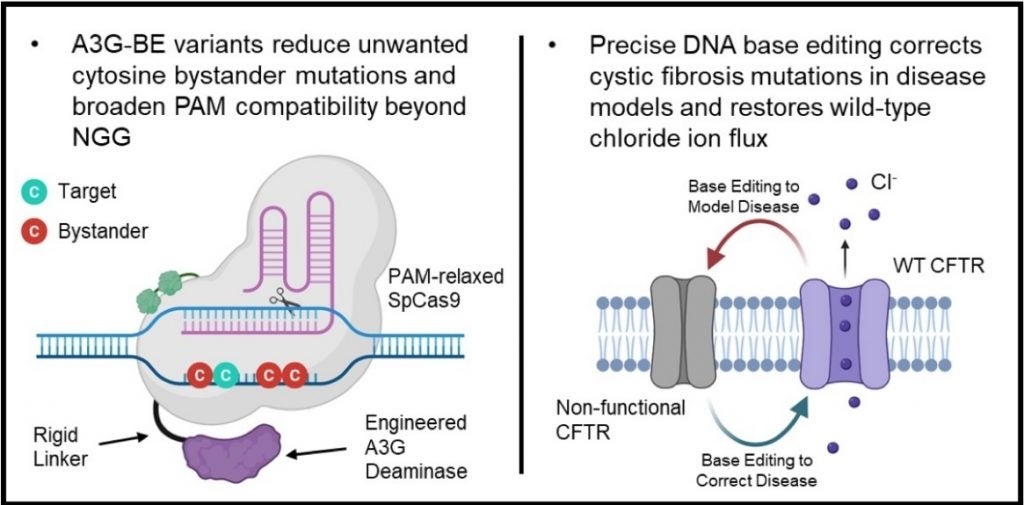
In order to change letters in DNA, base-pair editors combine two essential functions: one component that locates a specific sequence in the genome and another that modifies DNA. Those two parts are physically connected by a segment of molecules known as the “linker,” which dictates how freely the enzyme that carries out DNA editing moves at the target site.
The researchers also modified how strongly the base-pair editor interacts with DNA, weakening its tendency to act on neighboring letters.
In laboratory tests in human cells, the redesigned editor led to dramatic reductions in unintended bystander edits: The most accurate variant decreased bystander mutations by more than 80%, while retaining high levels of the original editor’s activity at the target site.
Cystic fibrosis is caused by mutations in a gene that controls how cells lining the lungs move salt and water in and out. When that process breaks down, thick mucus builds up in the lungs, making it hard to breathe and leaving patients vulnerable to repeated infections.
While drugs developed in recent years, such as Trikafta, have transformed life for many people with cystic fibrosis, the medicines must be taken daily and can carry staggering annual costs. Because many of the mutations that cause cystic fibrosis involve changes to just a single DNA letter, base-pair editors could, in principle, help treat the disease, even when Trikafta is ineffective — but only if they can avoid causing harmful off-target mutations.
The work remains at an early, preclinical stage. Still, at several cystic fibrosis-related genetic sites affecting a subset of patients, the refined editor reduced unintended bystander edits from as high as 50-60% to less than 1%, while largely preserving the desired DNA change.
By directly repairing the underlying genetic error, the method points toward the possibility of a long-lasting, potentially permanent treatment.
“The more precise we can make these tools, the greater their potential to change how we treat genetic disease with a high level of efficacy and safety,” Bao said.
Beyond cystic fibrosis, the refined base editor could help researchers tackle a wide range of genetic diseases caused by single-letter DNA changes. Because the tool allows scientists to introduce and correct specific mutations with much greater precision, it offers a powerful way to study how individual genetic variants affect disease and drug response.
That capability is especially valuable for rare mutations, which may affect only a small number of patients and are difficult to study through large clinical trials. By creating accurate cellular models of those mutations in the lab, researchers can test existing drugs, explore new therapeutic strategies and begin to identify which treatments are most likely to work for specific patients.
“That kind of insight is essential for moving toward more personalized approaches to treating genetic disease,” Gao said.
The research was supported by the National Institutes of Health (HL157714, HL173243, T32NS091006), the National Science Foundation (2143626, 2236662) and the Cystic Fibrosis Foundation (BAO19XX0, BAO19XX1). The content in this press release is solely the responsibility of the authors and does not necessarily represent the official views of funding entities.