HOUSTON – (Sept. 13, 2021) – Progress on treating Alzheimer’s disease has been frustratingly slow. A group of scientists in Houston suggest frustration at a very small scale may lead to a new path toward treatment.
Researchers at the University of Houston (UH) and at Rice University, associated with the Rice-based Center for Theoretical Biological Physics (CTBP), found through experiments and computations that amyloid beta peptides, small molecules that are abundant in the brain, go through several intermediate stages of frustration as they “dock and lock” to the tips of growing fibrils.
Folding proteins tend to look for the easiest way to get to their functional forms. Similarly, amyloid beta peptides look for the easiest way to bind to the tips of growing fibrils, but are sometimes held back -- or frustrated -- when the positive and negative forces between atoms don’t immediately align.
When they do finally align, the growing fibrils form the gummy plaques implicated in Alzheimer’s and other neurological diseases. New research in the Proceedings of the National Academy of Sciences shows that drugs might be developed to take advantage of the peptides’ frustrated intermediate states to stabilize the fibril tips and block further aggregation.
UH chemical and biomolecular engineer Peter Vekilov, said it wasn’t a stretch to look at amyloid beta fibril growth in his lab. “Previous studies that observed fibrilizing proteins with an atomic force microscope focused on more exotic behaviors because amyloid beta fibrils with a steady rate of growth are kind of boring,” he said.
“But I was fascinated because the correlation of the growth rate to the concentration of peptides in the solution carries loads of information,” Vekilov said. “It helps to measure the rate constant, a quantity which is easy to model.”
He said Rice physicist Peter Wolynes, whose lab specializes in building computer models of protein and chromosome folding, suggested that disrupting the steady growth with urea, known to denature (or unfold) proteins, might provide useful data about how amyloid fibrils form. It sure did.
“A weird thing happened,” Vekilov said. “Urea made the fibrils less stable, which meant the bonds between the molecules in the fibrils became less strong. But it also made them grow faster. This is a very serious contradiction, a violation of the empirical rules of chemistry.
“But there are empirical rules, and then there are fundamental laws,” he said. “We thought, this is trying to tell us something.”
Further experiments showed that urea “destabilized the wrong peptide bonds,” Vekilov said. “It made the fibril grow faster but also showed us the intermediate frustrated steps. The big thing is we now have evidence that at the end of the fibril there’s a crown of frustrated, disordered peptide chains trying to dock and lock, and these are druggable targets.
“It’s irrational to block every single peptide, because there are probably 100,000 times more of them than there are fibril tips,” he said. “The beauty of what we found is that the fibril tip is an Achilles’ heel of fibrilization, and all we have to do is block the complex at the tip.”
Wolynes noted there were signs of frustration in an earlier study that saw evidence of hitches in fibril aggregation. “Two things have emerged from the experiments,” he said. “One is that almost all of the kinetic models people use for amyloid beta growth are too simple. That’s not unexpected. The other is that denaturing changes the equilibrium, and it can also change the rate of folding in in ways that tell you where the transitions states appear.
“In the earlier paper on fibril nucleation, we remarked that it looked like there were some strange processes where the proteins had to backtrack from the transition state,” Wolynes said. “So Peter went and investigated this, and I think he is the first person to do so.”
He said having a way to halt fibrils from growing may help settle a long-standing disagreement among scientists over whether fibrils cause neurological disease or protect the brain from another suspect, particularly tangled tau proteins.
“Our idea is to poison the tip so that it can’t grow, rather than to destabilize the whole fibril,” Wolynes added. “This, of course, gets into the big argument over whether fibrils are good or bad.”
Computational models might show that arresting the fibrils could either stem the effects of Alzheimer’s or make it worse. Regardless, Wolynes said scientists will have a more definitive answer.
“To my mind, what’s interesting here is to provide a new target, and we will explore some possible drugs that could change the nature of the tip,” he said. “Either way, those molecules will provide interesting tools to understand how fibril growth happens.”
Graduate students Yuechuan (Alex) Xu of UH and Kaitlin Knapp of Rice are co-lead authors of the paper. Co-authors are Rice postdoctoral researcher Nicholas Schafer and visiting faculty member Aram Davtyan; UH alumnus Mohammad Safari, now a postdoctoral researcher at Princeton University; and Kyle Le, a UH alumnus and former Rice research intern. Vekilov is the John and Rebecca Moores Professor of Chemical and Biomolecular Engineering and a professor of chemistry at UH. Wolynes is the D.R. Bullard-Welch Foundation Professor of Science and a professor of chemistry, biosciences and physics and astronomy, and co-director of CTBP.
The National Science Foundation and NASA supported the research.
-30-
Read the abstract at https://doi.org/10.1073/pnas.2110995118.
This news release can be found online at https://news-network.rice.edu/news/2021/09/13/docking-peptides-slow-to-lock-open-possible-path-to-treat-alzheimers/
Related materials:
Computer simulations explore how Alzheimer’s starts: http://news.rice.edu/2016/10/03/computer-simulations-explore-how-alzheimers-disease-starts-2/
Tumbleweeds or fibrils: Tau proteins need to choose: http://news.rice.edu/2020/02/03/tumbleweeds-or-fibrils-tau-proteins-need-to-choose-2/
Vekilov Research Group: https://vekilovgroup.chee.uh.edu
Wolynes Research Group: https://wolynes.rice.edu
William A. Brookshire Department of Chemical and Biomolecular Engineering at UH: http://www.chee.uh.edu/
Cullen College of Engineering at UH: https://www.egr.uh.edu/
Center for Theoretical Biological Physics at Rice: https://ctbp.rice.edu
Wiess School of Natural Sciences at Rice: https://naturalsciences.rice.edu
Video:
Video produced from successive atomic force microscope images shows amyloid beta fibrils as they’re cut by the microscope tip and regrow. Researchers at the University of Houston and Rice University found evidence that amyloid beta peptides in a solution undergo several steps of frustration as they try to “dock and lock” to the growing fibrils. (Credit: Vekilov Research Group/University of Houston)
Images for download:
https://news-network.rice.edu/news/files/2021/08/0823_AMYLOID-1-WEB.jpg
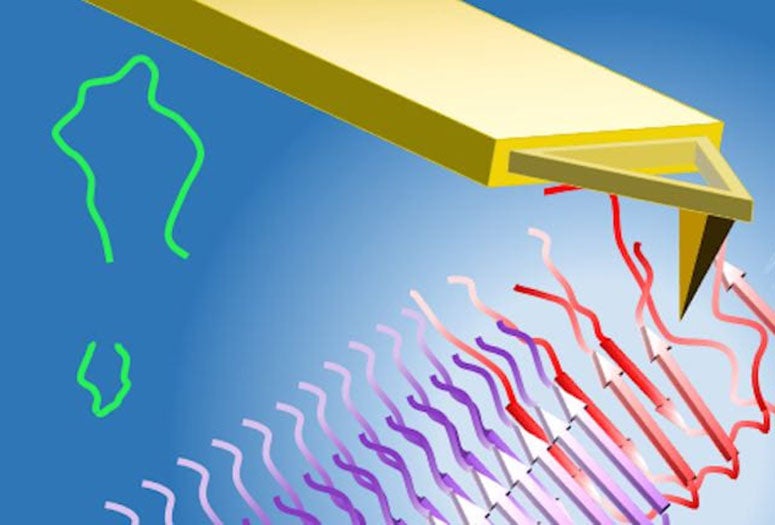
Using computational models and atomic force microscope experiments, researchers at the University of Houston and Rice University have identified a possible “Achilles’ heel” in the frustration of amyloid beta peptides as they dock to the fibrils that form plaques in patients with Alzheimer’s. The frustrated steps could open a window for drugs able to cap the fibril ends, preventing further aggregation. (Credit: Illustration by Yuechuan Xu/Peter Vekilov/University of Houston)
https://news-network.rice.edu/news/files/2021/08/0823_AMYLOID-2-WEB.jpg
A still from video of successive atomic force microscope images shows amyloid beta fibrils as they’re cut by the microscope tip and regrow. Researchers at the University of Houston and Rice University found evidence that amyloid beta peptides in a solution undergo several steps of frustration as they try to “dock and lock” to the growing fibrils. (Credit: Vekilov Research Group/University of Houston)
https://news-network.rice.edu/news/files/2021/08/0823_AMYLOID-3-WEB.jpg
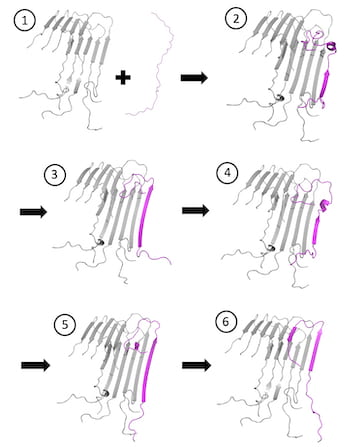
A sequence from computational models produced at Rice University shows how an amyloid beta peptide (magenta) docks and locks to an amyloid fibril. Simulations revealed the intermediate conformations in steps 2 through 5, indications of frustration in binding that may present new opportunities for treatment. (Credit: Illustration by Kaitlin Knapp/Rice University)
https://news-network.rice.edu/news/files/2021/08/0823_AMYLOID-5-WEB.jpg
Researchers at the University of Houston and Rice University have identified a possible “Achilles’ heel” in the frustration of amyloid beta peptides as they dock to the fibrils that form plaques in patients with Alzheimer’s disease. From left, Kyle Le, Kaitlin Knapp, Yuechuan (Alex) Xu, Peter Vekilov and Peter Wolynes. Missing from the photo are co-authors Nicholas Schafer, Aram Davtyan and Mohammad Safari. (Credit: Jeff Fitlow/Rice University)
https://news-network.rice.edu/news/files/2021/08/0823_AMYLOID-4-WEB.jpg
Graduate student Yuechuan (Alex) Xu, left, and Professor Peter Vekilov of the University of Houston with an atomic force microscope of the sort they used to capture the growth of amyloid beta fibrils implicated in Alzheimer’s disease. (Credit: Jeff Lautenberger/University of Houston)
About the University of Houston:
The University of Houston is a Carnegie-designated Tier One public research university recognized with a Phi Beta Kappa chapter for excellence in undergraduate education. UH serves the globally competitive Houston and Gulf Coast Region by providing world-class faculty, experiential learning and strategic industry partnerships. Located in the nation's fourth-largest city and one of the most ethnically and culturally diverse regions in the country, UH is a federally designated Hispanic- and Asian-American-Serving institution with enrollment of more than 47,000 students.
About Rice University:
Located on a 300-acre forested campus in Houston, Rice University is consistently ranked among the nation’s top 20 universities by U.S. News & World Report. Rice has highly respected schools of Architecture, Business, Continuing Studies, Engineering, Humanities, Music, Natural Sciences and Social Sciences and is home to the Baker Institute for Public Policy. With 3,978 undergraduates and 3,192 graduate students, Rice’s undergraduate student-to-faculty ratio is just under 6-to-1. Its residential college system builds close-knit communities and lifelong friendships, just one reason why Rice is ranked No. 1 for lots of race/class interaction and No. 1 for quality of life by the Princeton Review. Rice is also rated as a best value among private universities by Kiplinger’s Personal Finance.