Researchers co-led by a Rice University scientist have described for the first time specific genes and pathways in the brain associated with Alzheimer’s disease.
A paper published today by Rice computer scientist Vicky Yao and colleagues at Rockefeller and Princeton universities, among others, details molecular-level profiles of neurons in rodents similar to those in humans.
These details provide part of a framework to help understand complex interactions between amyloid beta proteins thought to be responsible for Alzheimer’s progression, aging and the neurodegeneration of vulnerable neurons that contain entangled tau proteins, according to the researchers.
The study appears in the Cell Press journal Neuron.
Yao, an assistant professor of computer science who joined Rice in 2019, began the study while a graduate student at Princeton, in collaboration with a group led by Nobel laureate Paul Greengard at Rockefeller. Greengard worked on the project until his death in 2019.
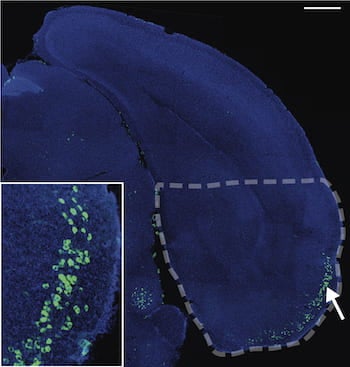
The study sought to understand why memory-forming neurons in the brain, particularly those in the entorhinal cortex (EC), are the first to die when Alzheimer’s takes hold. The researchers combined unique datasets collected by the Greengard lab with Yao’s innovative analysis to find the molecular drivers of the pathological cascade that appears to begin with amyloid aggregate formation throughout the brain and ends with tangled tau proteins in the neurons themselves.
In the process, Yao, co-lead author Jean-Pierre Roussarie of Rockefeller and their colleagues first demonstrated the strong similarities between humans and rodents in the conservation of cellular features in neurons that are either resistant or vulnerable to Alzheimer’s. That understanding raises confidence in using rodent models to develop therapies for humans.
They also pinpointed a gene involved in the constant microtubular remodeling of neuronal axons and describe its role in “selective vulnerability” to the disease. Yao said the plasticity of these vulnerable neurons in the EC may put them at risk but can also offer clues to new therapeutic targets.
“In most neurodegenerative diseases, the pattern in which cells die is very consistent,” she said. “The most vulnerable cells are always in the entorhinal cortex. We wondered if we could leverage that pattern to figure out what it is about these neurons that make them more vulnerable.”
For her part, Yao used computer models to map the interacting genes and predict what might be involved in neurodegeneration. That required establishing the strong molecular similarities between rodent and human neuron types so the models could be backed up with experiments.
The models showed the first evidence that tau-regulated microtubule processes center around the MAPT gene in vulnerable neurons, and may differ from those in resistant neurons. The network models also identified associations with aging and amyloid beta proteins.
They further pinpointed PTB, a splicing protein known to be dysregulated in Alzheimer’s, as the link between the two and an active contributor to tau aggregation in neurons in the EC.
“They constantly remodel themselves,” Yao said. “So the hypothesis is that if this tightly controlled process is thrown a little out of whack, the proportion of tau is different. The microtubules become hyperphosphorylated, which then leads the system to decide to kill it.”
Whether amyloid or tau ultimately causes the onset of Alzheimer’s has been the topic of debate among scientists for decades, she acknowledged.
“We didn't set out to try to link amyloid and tau, but as we were modeling it, when tau popped out, obviously there was a lot of excitement,” Yao said. “We’re finding the connection between them is subtle. There's a network of interactions and some disruption at a more nuanced level than expression, which is probably why it hasn't been identified to this kind of resolution before.”
While a treatment may not be imminent, she said better understanding of the network can only help.
“Once we figure out what makes neurons more vulnerable, that can lead to multiple avenues to decrease their vulnerability,” Yao said. “If we know what makes these neurons more and more vulnerable, up to cell death, we can look for a way to stop it.”
The research was supported by the National Institutes of Health (NIH), the Canadian Institute for Advanced Research, the U.S. Army Medical Research and Materiel Command, the National Institute on Aging, the Fisher Center for Alzheimer’s Disease Research, the JPB Foundation, the Cure Alzheimer’s Fund, the NIH Office of Research Infrastructure Programs and the National Institute of Neurological Disorders and Stroke.