David Ruth
713-348-6327
david@rice.edu
Jade Boyd
713-348-6778
jadeboyd@rice.edu
Caught in the act: Study probes evolution of California insect
First-of-its-kind study reveals parallel genomic changes during species formation
HOUSTON — (May 15, 2014) — A first-of-its-kind study this week suggests that the genomes of new species may evolve in a similar, repeatable fashion — even in cases where populations are evolving in parallel at separate locations. The research is featured on the cover of the May 16 issue of Science.

Researchers used a combination of ecological fieldwork and genomic assays to see how natural selection is playing out across the genome of Timema cristinae, a California stick insect that is evolving into two unique species. Credit: M. Muschick/University of Sheffield
A team of evolutionary biologists at Rice University, the University of Sheffield and eight other universities used a combination of ecological fieldwork and genomic assays to see how natural selection is playing out across the genome of a Southern California stick insect that is in the process of evolving into two unique species.
“Speciation is the evolutionary process that gives rise to new species, and it occurs when barriers prevent two groups of populations from exchanging genes,” said Rice co-author Scott Egan. “One way to study how speciation occurs is to look for examples where partial reproductive barriers exist but where genes are still exchanged.”
The stick insect Timema cristinae is one such example. Timema are closely related to “walking sticks,” plant-eating insects that look like twigs. Timema’s shape and color act as natural camouflage and help them avoid being eaten by predators, such as birds. More than a dozen unique species of Timema have evolved to feed on specific plants in California and northern Mexico.
One of these, T. cristinae, is found in two distinct varieties. One variety, or ecotype, feeds on the thin, needle-like leaves of a shrub called Adenastoma and features a distinct white stripe on its back that serves as camouflage. The other ecotype has no stripe and feeds on Ceanothus, a plant with wide green leaves where the stripe would stand out.
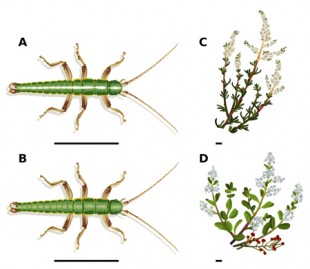
The species Timema cristinae has two distinct varieties, or ecotypes. One features a distinct white stripe on its back and feeds on the thin, needle-like leaves of a shrub called Adenastoma. The other phenotype has no stripe and feeds on Ceanothus. Credit: T. Farkas/University of Sheffield
“Populations of T. cristinae on the two host plants have evolved many differences in their physical form while still exchanging genes,” said Egan, a Huxley Faculty Fellow in Ecology and Evolutionary Biology at Rice. “These same populations have also evolved barriers to gene flow. We call this process ‘speciation with gene flow,’ and evolutionary biologists have long wondered if the genetic basis for this process is highly repeatable and if the genes involved are spread out across the whole genome or in a few discrete regions.”
To find out whether this was the case, Egan and a dozen co-authors led by University of Sheffield biologist Patrik Nosil conducted four years of detailed genomic and ecological tests. They first had to sequence the genome for T. cristinae and identify which portions of the genome corresponded to particular biological functions. They then collected about 160 T. cristinae from the wild. Samples were collected at several geographic locations and were equally split between the ecotypes on the two host plants.
“We resequenced the genome of each individual that we collected and looked at which genes were differentiated between populations adapted to different host plants,” Nosil said. “Because we also conducted an experiment in the field measuring evolution in real time, we gained information on how natural selection is pulling these populations apart.”

University of Sheffield graduate student Timothy Farkas collects a Timema specimen. Credit: P. Nosil/University of Sheffield
For example, the team found that many of the genetic differences were related to the biochemical function of metal ion binding, and metals are known to influence differences in pigmentation and mandible shape between the two T. cristinae ecotypes.
Previous ecological studies have shown that Timema do not migrate long distances. Because of this, the team expected to find evidence of localized gene flow among individuals collected at specific geographic locations. The genomic tests confirmed this, but they also revealed a pattern in the way that natural selection was playing out at each of the localities.
“In particular, we found that there were regions of the genome that exhibited significant differences between populations from host plant 1 and host plant 2, regardless of where the individuals were collected,” Nosil said.
That suggested that evolution might be occurring in the same repeatable fashion at each location. To further test this, the team devised an experiment to gather genomic data from individuals that were actively under selection.
“We took individuals from a mixed population of the striped versus the no-striped ecotype, and we transplanted them back into nature onto the two host plants in five different sets,” Egan said. “We allowed them to go an entire generation, and then we resampled those populations, resequenced the genome of the survivors and compared those to the ancestors that we started with a year before. We tried to match up the allele frequency shifts in this experiment with the genome-level differentiation that we observed in our genome-resequencing populations. And what we found was that many of the regions that were highly differentiated in nature were the exact same regions that were responding to our selection experiment.”
Egan said it was previously impossible to conduct this kind of study because of the expense of genomic tests. Though the genomes of many plant, animal and microbial species have been sequenced over the past decade, most of those are model organisms. Scientists use model organisms to study critical biological processes, but Egan said the study of nonmodel organisms is often the key to ecological questions, including those related to how the environment influences natural selection and speciation.
“The world of genomics is beginning to open up for people like me who don’t study model organisms,” Egan said. “This is allowing us to address, in new ways, questions that Darwin posed over 150 years ago.”
Study co-authors include Víctor Soria-Corrasco, Aaron Comeault and Timothy Farkas, all of the University of Sheffield; Zachariah Gompert of Utah State University; Thomas Parchman of the University of Nevada, Reno; Spencer Johnston of Texas A&M University; Alex Buerkle of the University of Wyoming; Jeffrey Feder of the University of Notre Dame; Jens Bast of the University of Göttingen, Germany; Tanja Schwander of the University of Lausanne, Switzerland; and Bernard Crespi of Simon Fraser University, Canada. The research was supported by the European Research Council, Utah State University and the Wellcome Trust Centre for Human Genetics.
-30-
High-resolution IMAGES are available for download at:
https://news2.rice.edu/files/2014/05/0515_STICK-Timema-orig.jpg
CAPTION: Researchers used a combination of ecological fieldwork and genomic assays to see how natural selection is playing out across the genome of Timema cristinae, a California stick insect that is evolving into two unique species.
CREDIT: M. Muschick/University of Sheffield
https://news2.rice.edu/files/2014/05/0515_STICK-diagram-lg.jpg
CAPTION: The species Timema cristinae has two distinct varieties, or ecotypes. One features a distinct white stripe on its back and feeds on the thin, needle-like leaves of a shrub called Adenastoma. The other phenotype has no stripe and feeds on Ceanothus.
CREDIT: T. Farkas/University of Sheffield
https://news2.rice.edu/files/2014/05/0515_STICK-Tim-lg.jpg
CAPTION: University of Sheffield graduate student Timothy Farkas collects a Timema specimen.
CREDIT: P. Nosil/University of Sheffield
https://news2.rice.edu/files/2014/05/0515_STICK-Timema2-lg.jpg
CAPTION: Timema cristinae
CREDIT: A. Comeault/University of Sheffield
https://news2.rice.edu/files/2014/05/0515_STICK-Egan-lg.jpg
CAPTION: Scott Egan
CREDIT: S. Egan/Rice University
Located on a 300-acre forested campus in Houston, Rice University is consistently ranked among the nation’s top 20 universities by U.S. News & World Report. Rice has highly respected schools of Architecture, Business, Continuing Studies, Engineering, Humanities, Music, Natural Sciences and Social Sciences and is home to the Baker Institute for Public Policy. With 3,920 undergraduates and 2,567 graduate students, Rice’s undergraduate student-to-faculty ratio is 6.3-to-1. Its residential college system builds close-knit communities and lifelong friendships, just one reason why Rice has been ranked No. 1 for best quality of life multiple times by the Princeton Review and No. 2 for “best value” among private universities by Kiplinger’s Personal Finance. To read “What they’re saying about Rice,” go here.


