Researchers work to untangle knots, slipknots in species separated by a billion years of evolution
Strings of all kinds, when jostled, wind up in knots. It turns out that happens even when the strings are long strands of molecules that make up proteins.
A new study by scientists at Rice University and elsewhere examines structures of proteins that not only twist and turn themselves into knots, but also form slipknots that, if anybody could actually see them, might look like shoelaces for cells.
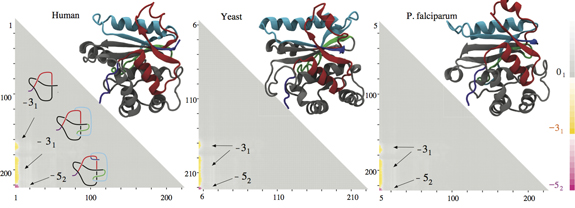
The molecular structures of the protein Ubiquitin C-terminal hydrolases from, left to right, humans, yeast and malaria-causing plasmodium falciparum cells form the same knotting motif, according to research by scientists associated with the Center for Theoretical Biological Physics. In all three cases, the proteins form 52 knots with nearly the same sizes and positions with respect to a linear map of their polypeptide chains. Courtesy of the Center for Theoretical Biological Physics
Proteins that serve the same essential functions in species separated by more than a billion years of evolution often display remarkable similarities. Joanna Sulkowska, a postdoctoral researcher at the Center for Theoretical Biological Physics (CTBP) at the University of California at San Diego, said these “strongly conserved” parts of proteins are especially common among those folds and hinges responsible for the knotted portions of a protein strand.
Sulkowska, co-first author of a new paper in the Proceedings of the National Academy of Sciences, works in the lab of her co-author, José Onuchic, Rice’s Harry C. and Olga K. Wiess Chair of Physics and a professor of physics and astronomy, chemistry, biochemistry and cell biology. Sulkowska expects to spend part of her year at CTBP when it moves its base of operation to Rice’s BioScience Research Collaborative this year.
She said slipknotted proteins, while rare, have been found in proteins that cross membrane barriers in cells. These transmembrane proteins stick through the cell membrane like pins in a pin cushion and help the cell sense and respond to its environment. “The slipknot is surprisingly conserved across many different families, from different species: bacteria, yeast and even human,” Sulkowska said. “They have really different evolutionary pathways, yet they conserve the same kind of motif. We think the slipknot stabilizes the location of the protein inside the membrane.”
Although a typical protein folds in a fraction of a second, researchers can see from simulations that knotted and slipknotted proteins would take longer to reach their folded structures than would unknotted proteins. Sulkowska said the extra effort to fold into knotted shapes must have a biological payoff or nature would have selected an easier path.
Finding the payoff is no easy task, but there are genomic clues. For instance, she said researchers suspect that “active sites” that control the folding pattern for knotted proteins often wind up inside the knotted structures after folding is complete. It’s possible, she said, that knotted proteins also have chaperone proteins that help the process along. Another mystery to be solved is how the body degrades knotted proteins; breaking down misfolded proteins is a normal function for healthy cells, and breakdowns in this process have been implicated in diseases like Alzheimer’s and Parkinsons.
Sulkowska, whose interest in knots extends to the macro realms of sailing and climbing, is sure there’s a good reason for all that she and Onuchic are seeing. “This is a new field, but we already know from experience how useful knots are,” she said. “They’re almost everywhere: in your shoes, in moving cargo, in physics as part of string theory. Now we hope to make this knowledge useful, maybe as a way to design new types of very stable proteins for disease treatment.
“Evolution didn’t redact these proteins,” she said. “They still fold, so they must have some function.”
Eric J. Rawdon of the University of St. Thomas, St. Paul, Minn., is co-first author. Co-authors include Kenneth Millett of the University of California at Santa Barbara and Andrzej Stasiak of the University of Lausanne, France.
The National Science Foundation, through CTBP, and the Swiss National Science Foundation supported the research.


Leave a Reply